Insulin 2.0? A new proteomics pipeline to discover novel endogenous bioactive peptides
Endogenous peptides are protein fragments that play important biological roles in the body. Famous examples include insulin that helps regulate blood sugar levels and endorphins that inhibit pain or promote euphoria. Some of these peptides, such as GLP-1, are ‘blockbuster drugs’ and have reached a commercial sale of more than $1 billion annually. We speculate that many such peptides are still to be discovered and therefore a great potential exists for both understanding basic biology better and at the same time identify new treatment options for diseases.
We have developed a streamlined analysis pipeline that lifts some of the haze surrounding the limited knowledge of the endogenous peptides that circulate within our bodies. Multiple challenges have so far restricted the number of endogenous peptides we can discover. A very big challenge originates from the fact that a major part of tissue and cells are made up of proteins. These are constantly being synthesized and degraded in body; hence a high number of detectable endogenous peptides are simply fragments or remnants of proteins that are in the process of being recycled. So how do we know if a peptide is simply a recycling product or a novel form of insulin?
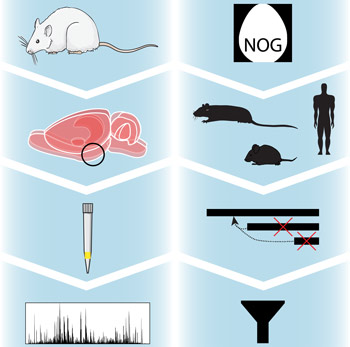
Figure: Our method relies on improvements in both sample handling and bioinformatics
Our new analytical method addresses this problem in three ways:
1) We inhibit recycling of proteins through application of chemical inhibitors and a physical trick, where heat is applied very rapidly. We show this has a benefit on the size of the peptide fragments that we find in rat brains.
2) We use our highly optimized and very fast peptide sequencing mass spectrometry platform to generate a large database of endogenous peptides extracted from hypothalamus, which comprises the biggest set of neuropeptides so far with more than 14,000 peptides found.
3) We accept that a large amount of our findings are not interesting and use a bioinformatics strategy to discriminate between peptides that have been specifically processed and those derived from unspecific recycling events. We demonstrate how this grouping works by rediscovering patterns of known processing patterns and find a link to site-specific peptide phosphorylation.
Phosphorylation is a process where peptides are modified chemically in-vivo by enzymes known as protein kinases. We show this post-translational modification is present at high levels on endogenous peptides and that it changes the properties and function of a biologically active peptide. In summary, we have developed a new discovery tool to identify a wealth of potentially new endogenous peptides and their regulatory modifications. Importantly, it can be used on any tissue or cell-type samples to address other unsolved questions in biology.
The work performed here has been performed in close collaboration with the pharmaceutical company Novo Nordisk.
Link to full article in Nature Communications: Analytic framework for peptidomics applied to large-scale neuropeptide identification

Novo Nordisk Foundation Center for Protein Research, University of Copenhagen is supported financially by the Novo Nordisk Foundation (Grant agreement NNF14CC0001)
