Visualizing phosphodiester-bond hydrolysis by an endonuclease
A press release from CNIO - the Spanish National Cancer Research Centre reveals how a group of scientists under Guillermo Montoya (formerly CNIO now Novo Nordisk Foundation Center for Protein Research, Faculty of Health and Medical Sciences at the University of Copenhagen) has developed a method for producing biological crystals. This enables the researchers to observe DNA double chain breaks for the first time. Moreover, the team has developed a computer simulation that makes this process visible to the human eye.
In the press release Professor Guillermo Montoya explains:
“We knew that enzymes, or proteins endonucleases are responsible for these double strand breaks but we didn’t know exactly how it worked until now,”
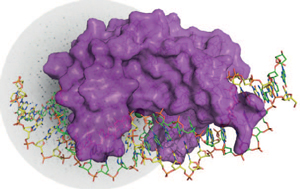
Crystal structure of the homing endonuclease (surface model in purple) in complex with its target DNA (sticks, each strand in green and yellow). The protein-DNA complex is depicted on top a X-ray diffraction pattern.
Authors:
Rafael Molina, Stefano Stella*, Pilar Redondo, Hansel Gomez, María José Marcaida, Modesto Orozco, Jesús Prieto & Guillermo Montoya*
*Novo Nordisk Foundation Center for Protein Research

