Researchers from the Nielsen group detail the human SUMO proteome
An emerging concept of modern biology relates to the systematic understanding of cellular networks co-regulated via post-translational modifications (PTMs). Hence, to better understand how cells function, researchers from the Nielsen group at CPR have mapped the human roadmap related to proteins modified with the small ubiquitin-like modifier (SUMO).
SUMOylation is a post-translational modification (PTM) analogous to ubiquitylation in terms of the enzymatic reaction cascade and types of enzymes used; however, instead of conjugating ubiquitin to target proteins SUMOylation involves the dynamic addition of SUMO.
SUMO has emerged as an essential modification that plays a crucial role in many biological processes in the nucleus of human cells, and is furthermore indispensable to all eukaryotic life. As a result, SUMOylation plays important roles in various human diseases including cancer, Alzheimer’s disease, Parkinson’s disease, Huntington’s disease, and ischemia. Consequently, the SUMO signaling pathway has emerged as a clinically relevant drug target; however, it has remained analytically challenging to obtain an accurate and global map of which proteins that specifically becomes modified by SUMO in human cells.
To address this challenge, researchers from the Nielsen group augmented a SUMO proteomics approach, hereby identifying 40,765 SUMO acceptor sites in 6,747 proteins, representing the first comprehensive map the human SUMOylome.
This map allowed the authors to better understand cellular processes surrounding SUMO and take the next step within the SUMO research field by establishing the distribution of SUMOylation across individual proteins. This provides researchers working on SUMO target proteins with valuable information regarding the specific abundances of each lysine residue modified by SUMO within the proteins of interest.
Moreover, using structural-predictive analyses the researchers find that SUMOylated lysine residues commonly reside in unstructured protein regions, in striking contrast to any other known lysine modification such as ubiquitylation, acetylation and methylation.
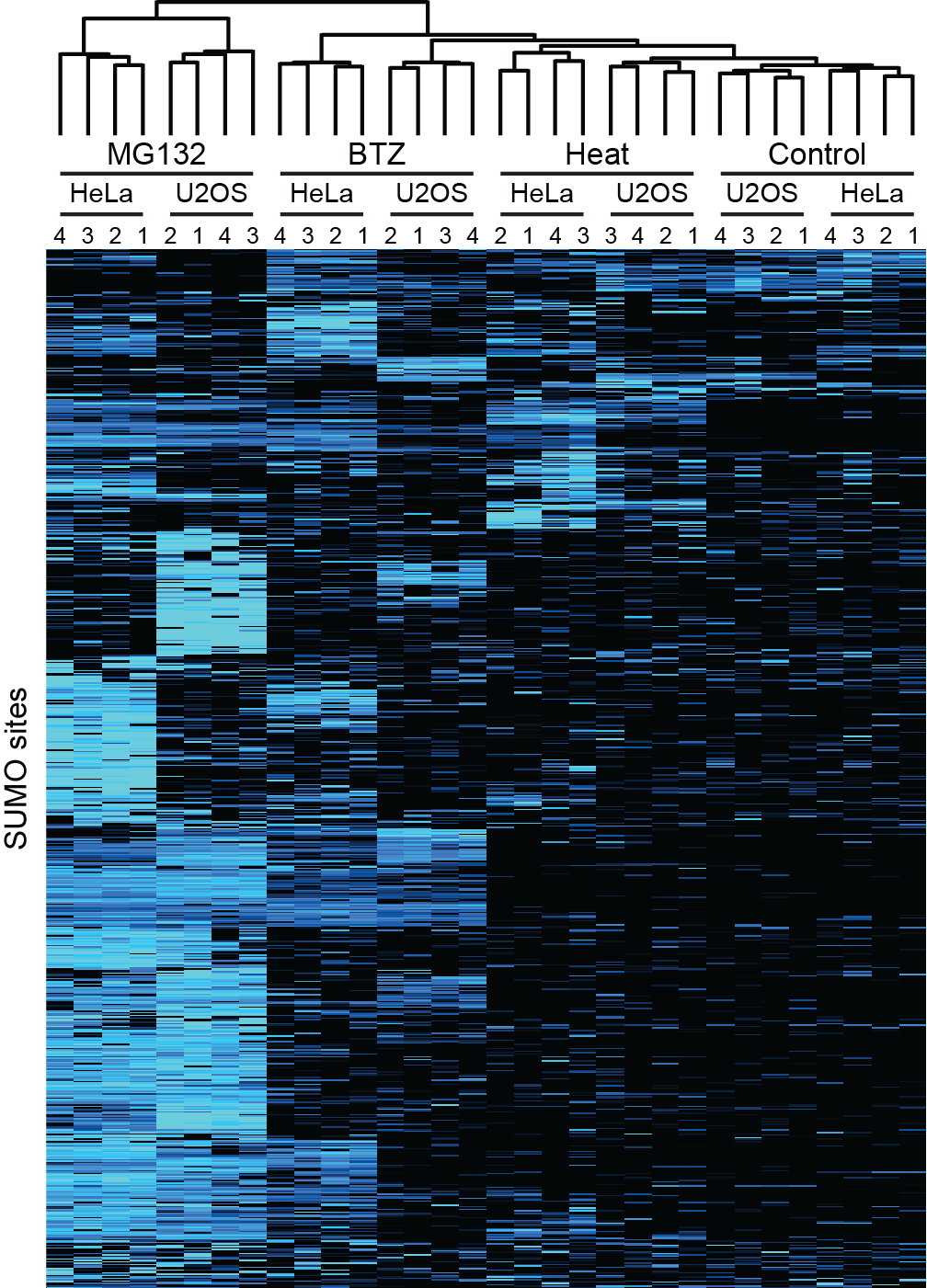
Figure 1: Quantitative regulation of SUMO-modification across cellular stress responses
Furthermore, the authors provide insight into SUMO dynamics in response to specific cellular stresses using quantitative mass spectrometry (Figure 1). To this end, the dynamic SUMOylation profiles where quantified for several relevant stresses, such as proteotoxic stress using the clinical proteasome inhibitor Bortezomib, and heat stress.
Besides describing proteins modified by SUMO, the researchers also investigated cellular cross-talk with other PTMs. They observed elaborate cross-modification between the various ubiquitin-like modifiers (UBLs), while discovering extensive co-modification between SUMO and phosphorylation (Figure 2).
Bioinformatics and proteomics analyses revealed SUMO modification events to be widely regulated in a manner dependent upon phosphorylation regulated by cyclin-dependent kinases (CDKs), demonstrating that phospho-dependent SUMOylation is involved in regulation of the cell cycle.
Collectively, the obtained data demonstrate that over 90% of all proteins integrated in nuclear protein complexes can become modified with SUMO, hereby detailing the SUMO proteome to unprecedented depth. The obtained data uncovers novel insights into the cellular functions of SUMO, and provides critical knowledge into human SUMO biology that undoubtedly will provide new opportunities for therapeutic strategies targeting the SUMOylation network in health and disease.
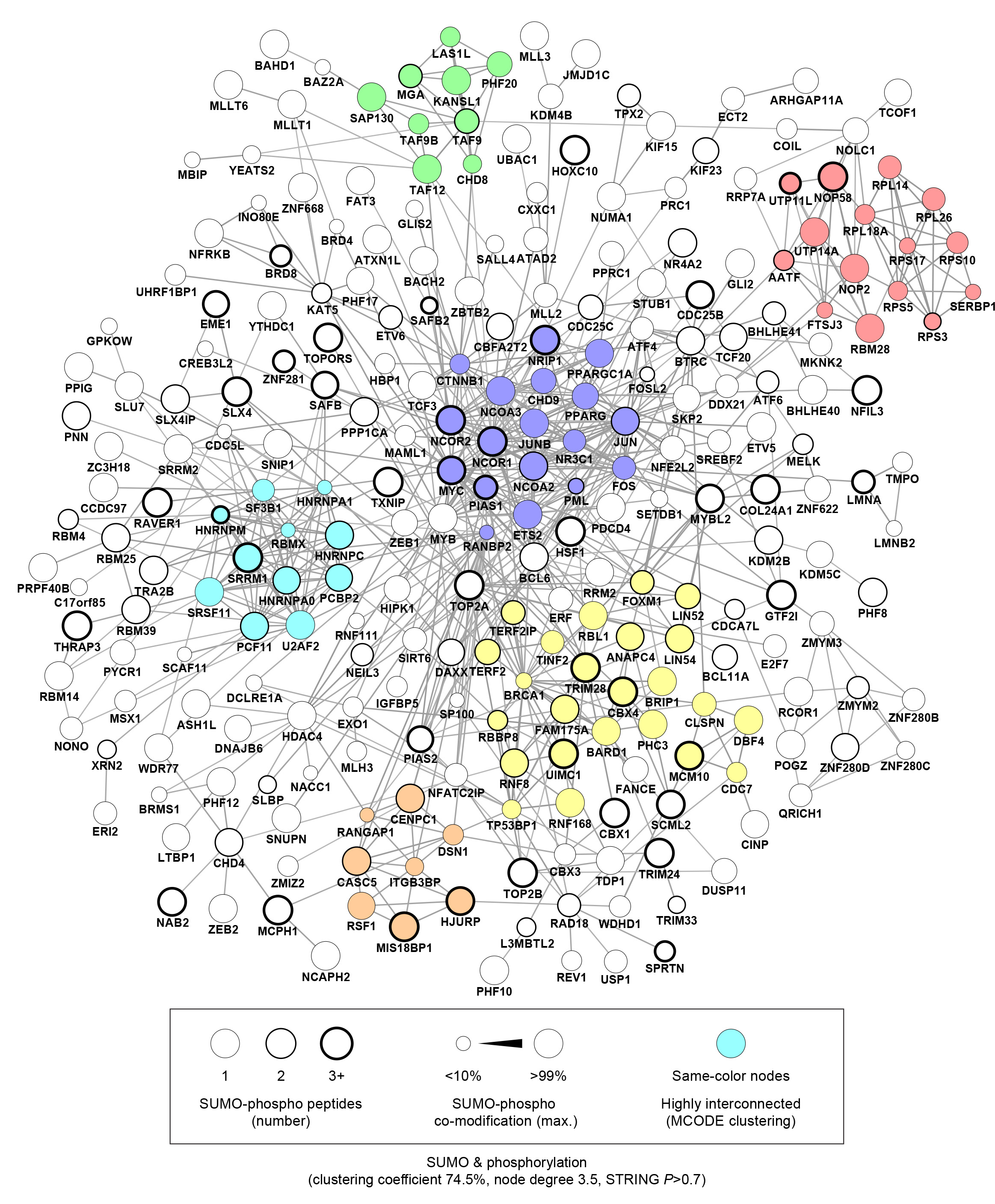 Figure 2: STRING visualization of co-regulated SUMO and phosphorylation networks.
Figure 2: STRING visualization of co-regulated SUMO and phosphorylation networks.
Authors from CPR:
Ivo Hendriks (Nielsen group)
David Lyon (Jensen group)
Clifford Young (Nielsen group)
Lars J. Jensen (Jensen group)
Michael L. Nielsen (Nielsen group)
Website of Nielsen Group at Novo Nordisk Foundation Center for Protein Research
Read the entire paper: 'Site-specific mapping of the human SUMO proteome reveals co-modification with phosphorylation'
Novo Nordisk Foundation Center for Protein Research, University of Copenhagen. The center is supported financially by the Novo Nordisk Foundation (Grant agreement NNF14CC0001)
