CPR researchers reveal new insights to the role of arginine methylation in disease occurrence & progression

The arginine methylation story published by the Nielsen group is featured on the cover of Science Signaling. The image shows a network of RNA-binding protein complexes that are regulated by arginine methylation
Researchers from the Nielsen group at Novo Nordisk Foundation Center for Protein Research (CPR) have, in collaboration with researchers from the Disease Systems Biology and Protein Signaling Programs also at CPR, published a comprehensive proteomics analysis on protein arginine methylation in human cells.
Posttranslational modifications, such as phosphorylation and ubiquitylation, regulate protein abundance, localization, interactions, and function. In this research article the Nielsen group investigated the landscape and functional roles of the emerging posttranslational modification arginine methylation.
The researchers found that arginine methylation is much more prevalent throughout the human proteome than previously anticipated, and they uncover that specific arginine methylation sites regulate the localization and function of splicing and RNA transport factors.
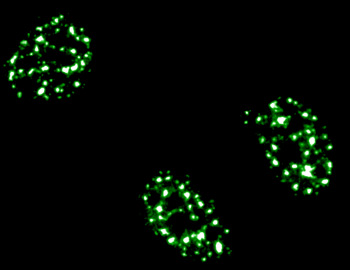
In the research article published by the Nielsen group, the authors investigate the PRMT-dependent regulation of SRSF2 using siRNA. A combined use of quantitative mass spectrometry and high-content imaging reveals a CARM1-specific regulation of SRSF2 into nuclear speckles
By using quantitative proteomics combined with RNA interference and high-throughput single-cell imaging the authors have revealed that arginine methylation specifically regulated two proteins involved in RNA processing and transport. The in depth analyses revealed that arginine methylation sites catalyzed by distinct arginine methyltransferases controlled the localization and RNA binding functions of the pre-mRNA splicing factor SRSF2 and the RNA-transporting activity of the protein HNRNPUL1.
On a broader level, the described data provide a rich resource for the future investigation of the function of protein arginine methylation. Moreover, detailed bioinformatic analyses indicate that sites modified by this posttranslational modification are hotspots for mutations in disease.
Principal investigator and Prof. Michael Lund Nielsen participated in Science Signaling podcast 30 August 2016 - Science Signaling Podcast for 30 August 2016: Human arginine methylome
